 Unfortunately, though, you will only have four features (unless your array is meant to be the other way around). There is no "correct" order or sign for the eigenvectors, so either is valid. They are very much related: The right singular vectors of A are. I use '~' to indicate that while they are "equal", the sign and order may vary. SVD is a decomposition for arbitrary-size matrices, while EIG applies only to square matrices. V is a matrix containing the eigenvectors, and D contains the eigenvalues. = svd(A) % Get the singular values of the original matrix = eig(X) % Get the eigenvectors and eigenvalues of the covariance matrix Check the following Matlab code: A = % A rectangular matrix This is good, because the singular values of a matrix are related to the eigenvalues of it's covariance matrix. The covariance matrix is not only square, it is symmetric. That is, if you have a matrix A, the covariance matrix is AA T. PCA actually uses the eigenvalues of the covariance matrix, not of the original matrix, and the covariance matrix is always square. Eigenvalues and eigenvectors only exist for square matrices, so there are no eigenvectors for your 150x4 matrix.Īll is not lost. However, I'm afraid I have bad news for you. See eigsvdgui in Numerical Computing with MATLAB or Cleve's Laboratory.The short (and useless) answer is that the eig(_) function gives you the eigenvectors and the eigenvalues. Cleve Molers Numerical Computing with MATLAB gives a nice overview of the differences between singular values and eigenvalues. The SVD computes the singular values of a matrix.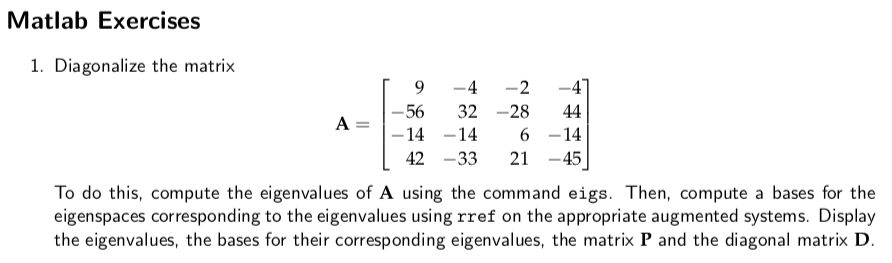 
These two sets of quantities are not the same. Is there a way to find out what the natural ordering of singular values would be The reason I ask is because the singular values correspond to dimensions associated with rows of the input matrix. Finally, you should not confuse singular values with eigenvalues. The resulting diagonal contains the singular values. MATLAB documentation of SVD states that the diagonal matrix returned has singular values in decreasing order. Now a two-sided QR iteration reduces the off diagonal to negligible size. Use a Householder operating from the left to zero a column and then another Householder operating from the right to zero most of a row. Make our test matrix rectangular by inserting a couple of rows of the identity matrix. It is related to the polar decomposition. It generalizes the eigendecomposition of a square normal matrix with an orthonormal eigenbasis to any matrix. The limit is the diagonal containing the eigenvalues. In linear algebra, the singular value decomposition ( SVD) is a factorization of a real or complex matrix. Now the QR iteration works on just two vectors, the diagonal and the off-diagonal. (The computation is done on half of the matrix, but we show the entire array.)īy symmetry the six Householders that zero the columns also zero the rows. The eigenvalues of a bump are a complex conjugate pair of eigenvalues of the input matrix. 1 The answer is to use a package, because mostly anything you try by hand will be woefully inefficient in python. 
This example has one bump in rows 3 and 4. This is good, because the singular values of a matrix are related to the eigenvalues of its covariance matrix. So the final matrix may have 2-by-2 bumps on the diagonal. The remaining subdiagonals require just one or two iterations each.Īll this is done with real arithmetic, although a real, nonsymmetric matrix may have complex eigenvalues. The next two rows require three iterations each. The element below the diagonal in the last row is the initial target it requires four iterations. The iteration count is shown in the title. The corresponding diagonal element is an eigenvalue.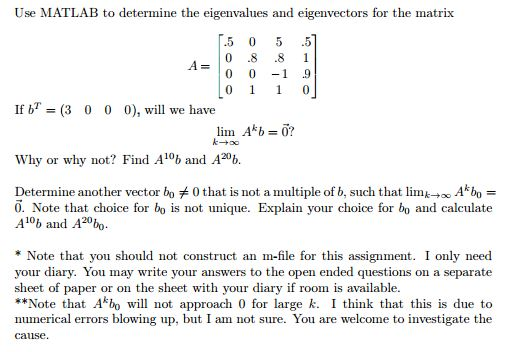 
Now the QR algorithm gradually reduces most subdiagonal elements to roundoff level, so they can be set to zero. The result is known as a Hessenberg matrix (don't let spell-checkers change that to Heisenberg matrix.) 
The initial reduction uses n-2 Householder similarites to introduce zeroes below the subdiagonal a column at a time. Here is a static picture of the starting matrix. The starting matrix for all three variants is based on flipping the Rosser matrix. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
